Acvtity stopped on Nov.1st, 2020
IBPM - CNR
Institute of Molecular Biology and Pathology - National Research Council, Italy
Nucleic Acids Laboratory
c/o Dept. Biology and Biotechnology, University «La Sapienza», Rome (Italy)
 |
MoMU
- The Molecular Microscopy Unit Acvtity stopped on Nov.1st, 2020 IBPM - CNR Institute of Molecular Biology and Pathology - National Research Council, Italy Nucleic Acids Laboratory c/o Dept. Biology and Biotechnology, University «La Sapienza», Rome (Italy) |
A selection of TEM and AFM images from the MoMU archives
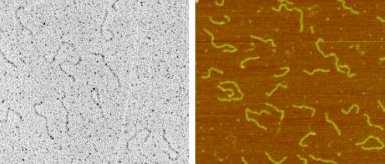 |
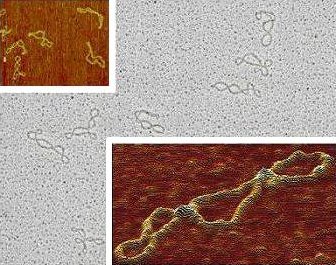
Bent DNA 520 bp restriction fragment carrying at one end the 211 bp bent region from Crithidia fasciculata kinetoplast DNA. The curvature is visualized on some molecules as a hairpin-like terminal fold. Preparation: adsorption on mica in the presence of magnesium ions. Left panel: TEM imaging upon rotary shadowing with platinum and carbon replica. Right panel: AFM imaging |
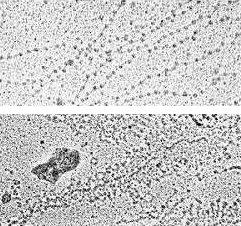 |
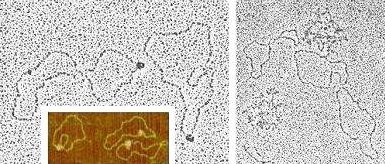
Chromatin Nuclear content of Xenopus laevis oocytes. Preparation: dispersal of chromatin in low ionic strength buffer, followed by fixation in sucrose/glutaraldehyde and adsorption on activated carbon films ('Miller spread'). TEM imaging upon rotary shadowing with platinum. Upper panel: beaded chromatin fibers (the beads, representing nucleosomes, have a diameter of about 120 Å). Lower panel: 'Christmas tree'-like arrangement of nascent ribosomal RNA transcripts along a portion (about 8 kb) of an rDNA repeat unit. |
 |
DNA supercoiling Supercoiled forms of plasmid pT7 (2.8 kb). Central panel: TEM imaging upon cytochrome-c/formamide spreading, uranyl acetate staining and rotary shadowing with platinum. Upper inset/lower inset (3D-view): AFM imaging upon adsorption on mica in the presence of magnesium ions. |
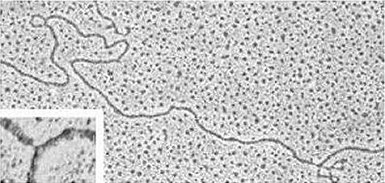 |
Heteroduplex. Replication
fork TEM imaging upon cytochrome-c/formamide spreading, uranyl acetate staining and rotary shadowing with platinum. Large panel: molecular hybrid (heteroduplex) between the antibiotic resistance transposons Tn21 and Tn1935. Tn1935 contains the entire sequence of Tn21, as well as two additional regions (0.97 kb and 2.7 kb, separated by a 7.8 kb stretch), corresponding to the genes for resistance to ampicillin and kanamycin, which appear as single-stranded DNA loops. Inset: replication fork in replicating DNA from Carassius auratus. The single stranded region at the fork point is about 120 Å long (35 bp). |
 |
In
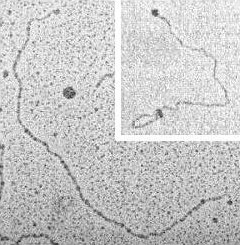
vitro transcription In vitro interaction between T7 RNA polymerase (about 100 kDa) and plasmid pT7 (2.8 kb) under conditions promoting (right panel) or inhibiting (left panel and inset) transcriptional activity. Preparation: adsorption of glutaraldehyde-fixed binary/ternary complexes on mica in the presence of magnesium ions. Left panel: TEM imaging upon rotary shadowing with platinum and carbon replica. Inset: AFM imaging. Right panel: TEM imaging as for left panel; RNA polymerase molecules are masked by the bush-like appearance of nascent transcripts. |
 |
DNA end-labelling In vitro interaction between terminally biotinilated, linearized plasmid pSP6 (3 kb) and streptavidin (about 65 kDa, left panel) or ferritin-conjugated avidin (about 900 kDa, inset). Left panel: TEM imaging upon BAC droplet-diffusion of binary complexes, uranyl acetate staining and rotary shadowing with tungsten. Inset: dark filed TEM imaging upon adsorption of binary complexes on mica in the presence of magnesium ions, rotary shadowing with tungsten and carbon replica. |
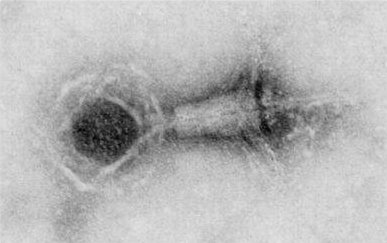 |
Bacteriophage TEM imaging of phage PBS1 upon negative staining with DMSO and uranyl acetate. The head has a diameter of about 0.120 µm. |
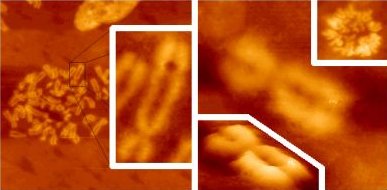 |
Metaphasic chromosomes AFM views of human metaphasic chromosomes from asynchronous fibroblast cultures. Left panel (47 µm x 47 µm field) and inset (9 µm x 18 µ field): air-imaging of the cell monolayer upon KCl hypotonic shock, methanol/acetic acid fixation and air drying. Central right panel (6 µm x 6 µm field): hypotonic shock/fixation as for left panel; imaging in liquid upon rehydration in PBS (lower inset shows a 3D-view of the same field). Upper right inset (20 µm x 20 µm): imaging in liquid of an unfixed metaphasic plate. |
[Top]
Last update: September 16th,
2022.
© 2006-2022 Gioacchino Micheli